This web page was produced as an assignment for Genetics 564 at UW-Madison Spring 2015.
What is the Tbx1 gene?
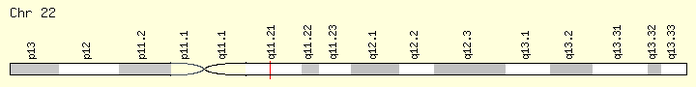
Genes are the basic instructions for life that coordinates what, how, when, and where cells need to function. It is estimated that approximately 20,000 to 25,000 genes make us, as humans, who we are (1). One of those 20,000 genes is Transcription Box 1, or Tbx1, which has been associated with 22q11.2 Deletion Syndrome (DS). Tbx1 is located in middle of the the deleted region on chromosome 22, as seen in Figure 1, and has been proven to be involved in the most distinct symptoms of 22q11.2 DS, including heart and craniofacial abnormalities (2). In 22q11.2 DS, one copy of Tbx1 is deleted while the other copy contains mutations or is not expressed due to a change in regulation of the gene, that causes the gene to loss its normal function (2). A wide range of mutations in Tbx1 has seen in patients with 22q11.2 DS, such as single nucleotide deletions causing a frameshift to a single nucleotide missense mutation (2), however, no genotype has been significantly associated with 22q11.2 DS (3).
HOMOLOGY
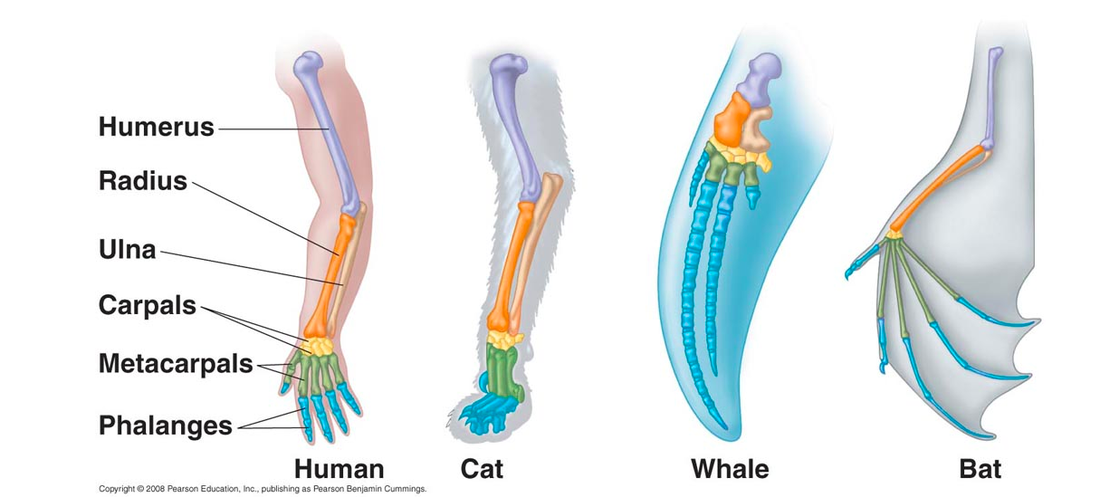
What is homology?Homology can be described as similarities between species due to the evolution from a common ancestor. Homologous structures basically look similar on the inside, but no longer have the same function or look the same on the outside between species. A great example of this can be seen by looking a the forelimb of different species. In figure 1, the structure of the limb remains constant even though it is well observed that a whale's flipper is used for swimming where as a bat's wing is used for flight (4). This can also be applied when looking at the structure and function of genes and proteins between species.
|
Homology of the Tbx1 gene
To find the homology of the Tbx1 gene, I first obtained the the gene accession number using ENTREZ by searching in the Nucleotide database that includes all DNA and RNA sequences and then filtered by RefSeq. By clicking on the GenBank tab, the human accession number was obtained first (NM_080647.1). The Tbx1 gene has three known and two predicted variants. I chose to precede with variant C due to findings in the literature and the length of the RNA sequence. This finding was confirmed by using OMIM and UNIPROT. Next, the location of the Tbx1 gene was found using the NCIB Map Viewer on Chromosome 22 on the q, or long arm, at position 11.21 (figure 1). This makes sense because the disease is named after the location of the deletion and Tbx1 is included in that region.
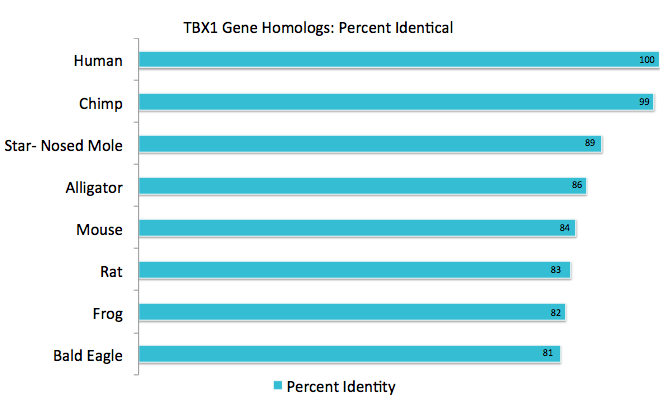
By using the human accession number, I was able to identify all of the homologs for Tbx1 by using BLAST and then aligning two sequences. This was also confirmed by using Homologene. Results are shown below in figure 3. Percent identity, or how much of the homologous sequence exactly matches the original, and E-value was noted. All E-values were close to zero meaning that findings are statically significant. A reciprocal BLAST was done to verify the findings by comparing the homologous organisms to the human Tbx1 gene.
By using the human accession number, I was able to identify all of the homologs for Tbx1 by using BLAST and then aligning two sequences. This was also confirmed by using Homologene. Results are shown below in figure 3. Percent identity, or how much of the homologous sequence exactly matches the original, and E-value was noted. All E-values were close to zero meaning that findings are statically significant. A reciprocal BLAST was done to verify the findings by comparing the homologous organisms to the human Tbx1 gene.
Analysis
The Tbx1 gene was found to be homologous in bilateria, or organisms with a front and back along with left and right sides, and remained fairly conserved in percent identity. This provides evidence that Tbx1 is essential for life. From earlier, I mentioned that Tbx1 is involved the formation of the heart and face. As seen in figure 3, I chose a wide variety of species to demonstrate how Tbx1 is working dynamically with other proteins to form a diverse range of facial structures. For example, think of a beak of an eagle in comparison to an snout of an alligator but yet they both contain similar Tbx1 gene sequences. It should be noted that more homologs were found with the Tbx1 protein than the gene.
Homologous DNA References:
Homo Sapiens (Human)
T-box 1, transcript variant C, mRNA
Accession Number: NM_080647
Length: 2082 bp
FASTA
Location: 22q11.21
T-box 1, transcript variant C, mRNA
Accession Number: NM_080647
Length: 2082 bp
FASTA
Location: 22q11.21
|
Alligator mississippiensis (American Alligator)
T-Box 1 Accession Number: XM_006270926 Max Identical: 86% E-Value: 0.0 FASTA |
|
Haliaeetus leucocephalus (Bald Eagle)
T-Box 1 Accession Number: XM_010581750 Max Identical: 81% E-Value: 0.0 FASTA |
|
Rattus norvegicus (Brown Rat)
T-Box 1 Accession Number: NM_001108322 Max Identical: 83% E-Value: 0.0 FASTA |
Pan troglodytes (Common Chimpanzee)
T-Box 1 Accession Number: XM_009437795 Max Identical: 99% E-Value: 0.0 FASTA |
|
Condylura cristata (Star-Nosed Mole)
T-Box 1 Accession Number: XM_004694935 Max Identical: 89% E-Value: 0.0 FASTA |
GENE ONTOLOGY
What is gene ontology?

Gene Ontology is the study of properties of gene products like proteins (5) . This is done by looking at the proteins biological process, cellular component, and molecular functions. A molecular function is defined as a single activity done at the molecular level. This is similar to biological process, however, a biological process describes a series of events to complete a molecular event. Finally, a cellular component is the location of the gene product in a cell (5).
The ontology of Tbx1 was found by using UNIPROT and then confirmed using QuickGO and the Gene Ontology Consortium. All biological processes are linked to AmiGO 2 for a brief description and accession number.
The ontology of Tbx1 was found by using UNIPROT and then confirmed using QuickGO and the Gene Ontology Consortium. All biological processes are linked to AmiGO 2 for a brief description and accession number.
Ontology of Tbx1
|
Biological Process:
Transcription Transcription regulation Heart development Pharyngeal system development Thyroid gland development Ear morphogenesis Face morphogenesis Cell differentiation |
Analysis
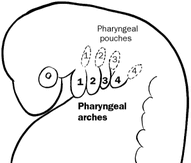
Tbx1 is involved in over 100 biological processes and reaffirms that Tbx1 is essential for life. From the list above, you can see that a majority of biological processes of Tbx1 related to a symptom of 22q11.2 DS. For example, Tbx1 is part of ear morphogenesis and pharyngeal system development and 22q11.2 DS can present with hearing loss and cleft palate. Since Tbx1 is a transcription factor it makes sense that it is localized where transcription takes place, in the nucleus. This also explains the DNA binding activity of Tbx1. Liao et al hypothesizes that Tbx1, like other transcription factors, function as dimers that bind to downstream targets, so when Tbx1 levels are low, less regulation of the downstream targets is taking place (7). These gene ontology findings support this hypothesis. The molecular function shows that Tbx1 is involved with dimerization activity, and the biological processes include positive regulation of other molecular events, such as cell proliferation.
References:
1. Genetic Home Reference (2015). http://ghr.nlm.nih.gov/handbook/basics/gene . Accessed March 22, 2015.
2. Online Mendelian Inheritance in Man (2012). http://omim.org/entry/602054 . Accessed March 22, 2015.
3. Guo, T., McGinn, D. M., Blonska, A., Shanske, A., Bassett, A., Chow, E., … the International Chromosome 22q11.2 Consortium. (2011). Genotype and cardiovascular phenotype correlations with TBX1 in 1,022 velo-cardio- facial/DiGeorge/22q11.2 deletion syndrome patients. Human Mutation, 32(11), 1278–1289. doi:10.1002/humu.21568
4. Encyclopedia Brittanica (2015). http://www.britannica.com/EBchecked/topic/270557/homology . Accessed March 22, 2015.
5. Gene Ontology Consortium. http://geneontology.org/page/ontology-documentation
6. UNIPROT. http://www.uniprot.org/uniprot/O43435. Accessed March 24, 2015.
7. Liao, J, et al. (2004). Full spectrum of malformations in velo-cardio-facial syndrome/ DiGeorge syndrome mouse model by altering Tbx1 dosage. Hum Mol Genetics. http://hmg.oxfordjournals.org.ezproxy.library.wisc.edu/content/13/15/1577
1. Genetic Home Reference (2015). http://ghr.nlm.nih.gov/handbook/basics/gene . Accessed March 22, 2015.
2. Online Mendelian Inheritance in Man (2012). http://omim.org/entry/602054 . Accessed March 22, 2015.
3. Guo, T., McGinn, D. M., Blonska, A., Shanske, A., Bassett, A., Chow, E., … the International Chromosome 22q11.2 Consortium. (2011). Genotype and cardiovascular phenotype correlations with TBX1 in 1,022 velo-cardio- facial/DiGeorge/22q11.2 deletion syndrome patients. Human Mutation, 32(11), 1278–1289. doi:10.1002/humu.21568
4. Encyclopedia Brittanica (2015). http://www.britannica.com/EBchecked/topic/270557/homology . Accessed March 22, 2015.
5. Gene Ontology Consortium. http://geneontology.org/page/ontology-documentation
6. UNIPROT. http://www.uniprot.org/uniprot/O43435. Accessed March 24, 2015.
7. Liao, J, et al. (2004). Full spectrum of malformations in velo-cardio-facial syndrome/ DiGeorge syndrome mouse model by altering Tbx1 dosage. Hum Mol Genetics. http://hmg.oxfordjournals.org.ezproxy.library.wisc.edu/content/13/15/1577