This web page was produced as an assignment for Genetics 564 at UW-Madison Spring 2015.
GENE EXPRESSION
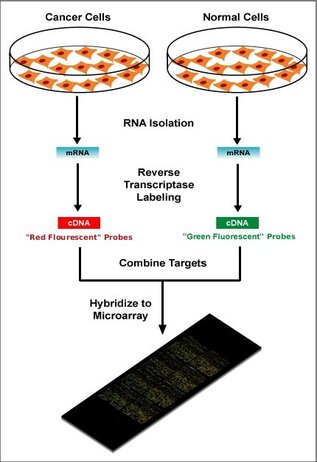
What is a microarray?A microarray is a way to study gene expression of the whole genome by measuring the amount of mRNA present in a sample (1). This can be used to compare gene expression between normal and disease states. Figure 4 shows how tumor, or disease, cell mRNA can be labeled with a color and the normal cell mRNA is labeled with a different color so you can differentiate between a change in gene expression by analyzing the colors on the microarray. An example analysis would be when a gene only appears green (normal cells) that means the tumor cells are no longer expression that gene and could be causing the disease state.
What is a GEO Profile?Gene Expression Omibus (GEO) is a database of gene expression profiles. A GEO profile consists of the gene expression results of an entire genome in specific samples between a normal verse disease state (1). GEO also has a cluster analysis program through heat maps. In heat maps, normal and disease gene expression can be studied by labeling them in different colors making it clear what is being expressed in a given state (2). GEO was used to search for gene expression experiments pertaining to expressions levels in organisms with 22q11.2 DS compared to normal. BioCarta was also used to determine if Tbx1 is part of a known pathway.
|
Analysis
No experiments studying 22q11.2 DS or a known pathway for Tbx1 was found. This would be an experiment worth doing since it is thought that severity in 22q11.2 DS is caused by abnormal Tbx1 levels (7). This would allow a researcher not only to compare how Tbx1 levels are changes between 22q11.2 DS phenotypes but also what other genes are being affected by this deletion syndrome. This can be done by looking at specific tissues that are known to be affected by 22q11.2 DS, like the ears or palate, at different stages of development. The information provided by this gene expression analysis could reveal evidence for a developmental pathway that includes Tbx1.
MODEL ORGANISMS
Why use model organisms to study disease? |
What is RNA Interference (RNAi)? |
|
The NIH defines a model organism as "an animal, plant or microbe that can be used to study certain biological processes." Due to ethical constrains, we cannot do certain experiments on humans, however, research has shown us that human diseases can be recreated in other organisms that are easier to use (4). There are still many considerations to take into account when choosing a model organism, some examples being time it takes to reproduce, life span, cost, maintenance level, and if the biological process happens in the organism.
|
RNAi is a method commonly used today to reduce function of a gene by targeting a specific gene mRNA for degradation. This is done by the use of small interfering RNAs (siRNA) or micro RNAs (miRNA). Both target a specific mRNA and from a degradation complex (RISC). Since the mRNA is being degraded before the it can be translated into a protein, gene expression is reduced (3). That being said, this method is known as a knock down instead of a knock out because some mRNA will escape being degraded and will become a functional protein, so there will just be less protein to be doing a given function. In a knock out scenario, the gene is gone so there is no chance that the given protein will be made.
|
TBX1 Mutant Phenotypes
Mutant phenotypes were found by searching human Tbx1 homolog databases: WormBase, FlyBase, MGI, and ZFIN.
C.Elegans (5)
Protein: mls-1
Phenotypes: None reported
GO terms:
biological processes- muscle cell fate specification, post embryonic development, and regulation of transcription
cellular component- nucleus
molecular function- transcription factor
Methods: RNAi was primarily used to reduce function of mls-1
Phenotypes: None reported
GO terms:
biological processes- muscle cell fate specification, post embryonic development, and regulation of transcription
cellular component- nucleus
molecular function- transcription factor
Methods: RNAi was primarily used to reduce function of mls-1
Drosophila (6)
Protein: org-1
Phenotypes: Manifested as adult external thorax, embryonic/larval circulatory system, embryonic/larval dorsal vessel, developing material anatomically entity, A2-7 ventral transverse muscle 1, unguis, embryonic heart visceral muscle, retina, circulatory systems, somatic precursor cell; Class: lethal
Go terms:
biological processes- positive regulation of transcription of RNA polymerase II promoter and somatic muscles development
cellular component- nucleus
molecular function- transcription factor
Methods: RNAi was primary used to reduce function of org-1 to get phenotype listed above
Phenotypes: Manifested as adult external thorax, embryonic/larval circulatory system, embryonic/larval dorsal vessel, developing material anatomically entity, A2-7 ventral transverse muscle 1, unguis, embryonic heart visceral muscle, retina, circulatory systems, somatic precursor cell; Class: lethal
Go terms:
biological processes- positive regulation of transcription of RNA polymerase II promoter and somatic muscles development
cellular component- nucleus
molecular function- transcription factor
Methods: RNAi was primary used to reduce function of org-1 to get phenotype listed above
Zebrafish (7)

Protein: tbx-1
Phenotypes: 70 phenotypes have been observed including abnormalities in the thymus, pharyngeal system, heart and ear
Go terms:
biological processes- involved in 29 processes including heart looping, pharyngeal system development, neural crest cell development, and ear morphogenesis
cellular component- nucleus
molecular function- sequence specific DNA binding transcription factor activity and protein homodimerization activity
Methods: tbx1 translation blocking morpholinos
Phenotypes: 70 phenotypes have been observed including abnormalities in the thymus, pharyngeal system, heart and ear
Go terms:
biological processes- involved in 29 processes including heart looping, pharyngeal system development, neural crest cell development, and ear morphogenesis
cellular component- nucleus
molecular function- sequence specific DNA binding transcription factor activity and protein homodimerization activity
Methods: tbx1 translation blocking morpholinos
Mouse (8)

Protein: tbx-1
Phenotypes: Homozygous null mice display neonatal lethality, persistent truncus arteriosis, abnormal aortic arch, abnormal inner, middle, and outer ear morphology, abnormal lymphangiogenesis, and abnormal cranial base morphology. Heterozygous null mice display abnormal fourth aortic arch arteries.
Go terms:
biological processes- involved in 113 processes including thymus and thyroid development, heart development, positive regulation mechanisms, and pharyngeal system development
cellular component-nucleus
molecular function-sequence specific DNA binding transcription factor activity and protein homodimerization activity
Methods: gene expression assays
Phenotypes: Homozygous null mice display neonatal lethality, persistent truncus arteriosis, abnormal aortic arch, abnormal inner, middle, and outer ear morphology, abnormal lymphangiogenesis, and abnormal cranial base morphology. Heterozygous null mice display abnormal fourth aortic arch arteries.
Go terms:
biological processes- involved in 113 processes including thymus and thyroid development, heart development, positive regulation mechanisms, and pharyngeal system development
cellular component-nucleus
molecular function-sequence specific DNA binding transcription factor activity and protein homodimerization activity
Methods: gene expression assays
Analysis
Through analysis of these results, it is clear that the choice is between zebrafish and mice due to shared key phenotypes and GO biological processes with humans. A lot of research for 22q11.2 DS has been done in these two model organisms for these reasons, however, I believe a zebrafish model best suits my goals outlined in my specific aims. Even though Tbx1 in zebrafish is not as highly conserved as other species, zebrafish have a well established molecular (Van Gogh) and behavioral model that I will utilize in my specific aims.
CHEMICAL GENETICS
What is chemical genetics?
Chemical genetics utilizes small molecules that bind to proteins in a way that changes the protein's function. This is another way of studying the function of a gene without using the traditional knockout method that can eventually lead to development of a new drug (9).
Both PubChem and ChemBank were used to search for small molecules that interaction with Tbx1 or similar proteins.
Model organism databases where also examined for use of chemical genetics.
Both PubChem and ChemBank were used to search for small molecules that interaction with Tbx1 or similar proteins.
Model organism databases where also examined for use of chemical genetics.
Tbx1 findings
No results were found using PubChem or ChemBank, however, RGD stated that 6-propyl-2-thiouracil, bisphenol A, and C60 fullerene were found to interact with Tbx1 in rats (10).
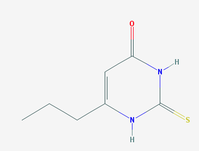
Chemical: 6-propyl-2-thiouracil
Methods: Using microarry analysis, Shiraki et al demonstrated that rats exposed to 6-propyl-2-thiouracil in the drinking water showed expression changes in genes participating in neuron differentiation and development, cell migration, synaptic function, and axon formation (11).
Results: Tbx1 mRNA decreases as 6-propyl-2-thiouracil increased (10)
Methods: Using microarry analysis, Shiraki et al demonstrated that rats exposed to 6-propyl-2-thiouracil in the drinking water showed expression changes in genes participating in neuron differentiation and development, cell migration, synaptic function, and axon formation (11).
Results: Tbx1 mRNA decreases as 6-propyl-2-thiouracil increased (10)
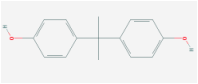
Chemical: Bisphenol A
Methods: Using microarray analysis and immunoctyochemistry, Ali et al demonstrated that bisphenol A interfered with meiosis and low doses altered expression of genes involved in the reproductive system (12).
Results: Tbx1 mRNA decreases in the presence of bisphenol A (10)
Methods: Using microarray analysis and immunoctyochemistry, Ali et al demonstrated that bisphenol A interfered with meiosis and low doses altered expression of genes involved in the reproductive system (12).
Results: Tbx1 mRNA decreases in the presence of bisphenol A (10)
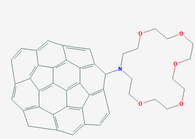
Chemical: C60 fullerene
Methods: Fujtak et al used gene expression profiles to examine the affects of exposure to C60 fullerene on the pulmonary system including genes involved in inflammatory, cell death, and immune responses (13).
Results: C60 fullerene Increases levels of Tbx1 mRNA (10)
Methods: Fujtak et al used gene expression profiles to examine the affects of exposure to C60 fullerene on the pulmonary system including genes involved in inflammatory, cell death, and immune responses (13).
Results: C60 fullerene Increases levels of Tbx1 mRNA (10)
Analysis
In these experiments, the researchers were looking at biological processes that Tbx1 is involved in, such as cell cycle regulation and differentiation. So it is no surprise to see expression changes in the presence of these chemical compounds. Most interesting to me is how 6-propyl-2-thiouracil directly alters Tbx1 expression specifically in the neuron. This provides a brain specific way to knockdown Tbx1 levels to observe the role of Tbx1 in neurogenesis. This can be used to study how decreases in Tbx1 levels in the brain affect specific 22q11.2 DS symptoms related to brain development and function, such as autism.
A last consideration for using potential drug intervention for 22q11.2 DS is that by the time a child is diagnosed there are already developmental abnormalities that cannot be reversed thus the drug rendering unless to the major causes of death in 22q11.2 DS patients. Due to this reason, chemical genetics will be limited to just a research tool to study 22q11.2 DS.
A last consideration for using potential drug intervention for 22q11.2 DS is that by the time a child is diagnosed there are already developmental abnormalities that cannot be reversed thus the drug rendering unless to the major causes of death in 22q11.2 DS patients. Due to this reason, chemical genetics will be limited to just a research tool to study 22q11.2 DS.
References:
1. About GEO Profiles (2014). http://www.ncbi.nlm.nih.gov/geo/info/profiles.html. Accessed March 24, 2014.
2. GEO Dataset Cluster Analysis (2014). http://www.ncbi.nlm.nih.gov/geo/info/cluster.html. Accessed March 24, 2015.
3. NIH: RNAi. (2014) http://www.ncats.nih.gov/research/reengineering/ncgc/rnai/rnai.html. Accessed March 24, 2015.
4. Using Model Organisms to Study Health and Disease. (2012) http://www.nigms.nih.gov/Education/Pages/modelorg_factsheet.aspx. Accessed March 25, 2015.
5. WormBase. http://www.wormbase.org/#01-23-6. Accessed March 24, 2015.
6. FlyBase. http://flybase.org/. Accessed March 24, 2015.
7. The Zebrafish Model Organism Database. http://zfin.org/cgi-bin/webdriver?MIval=aa-ZDB_home.apg. Accessed March 24, 2015.
8. Mouse Genome Informatics. http://www.informatics.jax.org/. Accessed March 24, 2015.
9. Using Small Molecules to Modulate a Protein. (2015) http://www.hhmi.org/biointeractive/small-molecule-microarrays. Accessed March 25, 2015.
10. RGD. http://www.rgd.mcw.edu. Accessed March 24, 2015.
11. Shiraki, A et al. (2014). Expression alternations of genes on both neuronal and glial development in rats after developmental exposure to 6-propyl-2-thiouracil. ELSERIER, Volume 228, Issue 3, 225-234. http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0378427414001775
12. Ali, S et al. (2014). Exposure to low-dose biosphenol A impaired meiosis in the rate somniferous tubules culture model: a physiotoxicogenomic approach. PLoS One, journal.prone.0106245, eCollection 2014. http://www.ncbi.nlm.nih.gov/pubmed?cmd=Search&term=25181051
13. Fujta, K et al. (2009). Gene expression profiles in rat lung after inhalation exposure to C60 fullerene particles. ELSEVIER, Volume 258, Issue 1, 47-55. http://www.sciencedirect.com/science/article/pii/S0300483X09000262
1. About GEO Profiles (2014). http://www.ncbi.nlm.nih.gov/geo/info/profiles.html. Accessed March 24, 2014.
2. GEO Dataset Cluster Analysis (2014). http://www.ncbi.nlm.nih.gov/geo/info/cluster.html. Accessed March 24, 2015.
3. NIH: RNAi. (2014) http://www.ncats.nih.gov/research/reengineering/ncgc/rnai/rnai.html. Accessed March 24, 2015.
4. Using Model Organisms to Study Health and Disease. (2012) http://www.nigms.nih.gov/Education/Pages/modelorg_factsheet.aspx. Accessed March 25, 2015.
5. WormBase. http://www.wormbase.org/#01-23-6. Accessed March 24, 2015.
6. FlyBase. http://flybase.org/. Accessed March 24, 2015.
7. The Zebrafish Model Organism Database. http://zfin.org/cgi-bin/webdriver?MIval=aa-ZDB_home.apg. Accessed March 24, 2015.
8. Mouse Genome Informatics. http://www.informatics.jax.org/. Accessed March 24, 2015.
9. Using Small Molecules to Modulate a Protein. (2015) http://www.hhmi.org/biointeractive/small-molecule-microarrays. Accessed March 25, 2015.
10. RGD. http://www.rgd.mcw.edu. Accessed March 24, 2015.
11. Shiraki, A et al. (2014). Expression alternations of genes on both neuronal and glial development in rats after developmental exposure to 6-propyl-2-thiouracil. ELSERIER, Volume 228, Issue 3, 225-234. http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0378427414001775
12. Ali, S et al. (2014). Exposure to low-dose biosphenol A impaired meiosis in the rate somniferous tubules culture model: a physiotoxicogenomic approach. PLoS One, journal.prone.0106245, eCollection 2014. http://www.ncbi.nlm.nih.gov/pubmed?cmd=Search&term=25181051
13. Fujta, K et al. (2009). Gene expression profiles in rat lung after inhalation exposure to C60 fullerene particles. ELSEVIER, Volume 258, Issue 1, 47-55. http://www.sciencedirect.com/science/article/pii/S0300483X09000262